Transection of vessels in epiphyseal cartilage canals leads to osteochondrosis and osteochondrosis dissecans in the femoro-patellar joint of foals; a potential model of juvenile osteochondritis dissecans.
"Ten Norwegian Fjord Pony foals were operated at the age of 13-15 days. Two vessels supplying the epiphyseal growth cartilage of the lateral trochlear ridge of the left distal femur were transected in each foal. Follow-up examination was carried out from 1-49 days post-operatively and included plain radiography, macroscopic and histological examination.
Transection[transection refers to division in half] of blood vessels within epiphyseal cartilage canals resulted in necrosis of vessels and chondrocytes, i.e. ischaemic chondronecrosis, in foals. Areas of ischaemic chondronecrosis were associated with a focal delay in enchondral ossification (osteochondrosis) in foals examined 21 days or more after transection, and pathological cartilage fracture (osteochondrosis dissecans) in one foal examined 42 days after transection."
"In epiphyseal growth cartilage, cartilage canals are regularly distributed, blind-ending tubular spaces that are present during the early phases of growth . An arteriole, its capillary bed and one or more venules course into and out of the cartilage through a single canal, and therefore represent true end arteries (“glomeruli”) . When they are no longer needed, cartilage canals regress by a physiological process known as chondrification, during which vessels disappear and perivascular mesenchymal cells within the canal differentiate into chondrocytes to fill the canal lumen with cartilage. Cartilage canals are also incorporated into the advancing ossification front during growth, and in piglets and foals, it has been suggested that the vessels are particularly vulnerable to failure during the incorporation process"
"“Programmed” chondrification when cartilage canals are no longer needed occurs within regions with thin cartilage before regions with thick cartilage"
Growing Taller: How Mesenchymal Stem Cells, Microfractures, Hydrostatic Pressure, and Periosteum makes increasing height possible
Friday, February 27, 2009
Tuesday, February 24, 2009
LCORL
Expression Levels of LCORL Are Associated with Body Size in Horses.
"Genome-wide association analyses revealed the highest associated quantitative trait locus for height at the withers on horse chromosome (ECA) 3 upstream of the candidate gene LCORL. The highly associated single nucleotide polymorphism BIEC2-808543 (-log(10)P = 8.3) and the adjacent gene LCORL [is] the most promising candidate for body size. We investigated the relative expression levels of LCORL and its two neighbouring genes NCAPG and DCAF16 using quantitative real-time PCR (RT-qPCR). We could demonstrate a significant association of the relative LCORL expression levels with the size of the horses and the BIEC2-808543 genotypes within and across horse breeds. In heterozygous C/T-horses expression levels of LCORL were significantly decreased by 40% and in homozygous C/C-horses by 56% relative to the smaller T/T-horses. Bioinformatic analyses indicated that this SNP T>C mutation is disrupting a putative binding site of the transcription factor TFIID which is important for the transcription process of genes involved in skeletal bone development. Thus, our findings suggest that expression levels of LCORL play a key role for body size within and across horse breeds and regulation of the expression of LCORL is associated with genetic variants of BIEC2-808543."
" the C allele of BIEC2-808543 is presumably the reason for the reduced expression of LCORL in larger sized horses. LCORL, also known as the Mblk1-related protein, shows characteristic motifs of transcription factors and analyses with mouse tissues indicate that it is able to activate transcription."<-So less LCORL equals greater height.
"In human genome-wide scans for adult stature evidences LCORL to be associated with trunk length and hip axis length"
"the correlation between growth of body and hair could be shown in various studies. The Rothmund-Thomson syndrome in human for example is characterized by severe dwarfism combined with an abnormal hair growth. Studies in mice revealed growth retardation in hair length and a retarded rate of body growth caused by the supply of high concentrations of the epidermal growth factor (EGF)"
So we can look at hair growth to find height genes.
"The candidate genes NCAPG and DCAF16 could be eliminated as candidates [for human height], whereas HMGA2 (high mobility group AT-hook 2, ECA6), ZFAT (zinc finger and AT hook domain containing, ECA9), LASP1 (LIM and SH3 protein 1, ECA11) could also possibly involved."
Four loci explain 83% of size variation in the horse.
"Unlike humans, which are naturally reproducing and possess many genetic variants with weak effects on size, we show that horses, like other domestic mammals, carry just a small number of size loci with alleles of large effect. Furthermore, three of our horse size loci contain the LCORL, HMGA2 and ZFAT genes that have previously been found to control human height. The LCORL/NCAPG locus is also implicated in cattle growth and HMGA2 is associated with dog size. Extreme size diversification is a hallmark of domestication."
"the immediately adjacent gene [to LCORL], NCAPG, has been implicated in prenatal growth"
"HMGA2 is an architectural transcription factor that regulates gene expression and directs cellular growth, proliferation and differentiation"
"1% of all human genes are now implicated in contributing to size variation"
Reaching new heights: insights into the genetics of human stature
"The height of an individual is the result of many growth and development processes and greater stature is not just a result of increasing bone length; tissue and organ sizes are usually also proportionately increased."
HMGA2 and GDF5 are two genes listed as highly influencing height.
LCORL along with c4orf30 are in the same gene loci as NCAPG.
"Genome-wide association analyses revealed the highest associated quantitative trait locus for height at the withers on horse chromosome (ECA) 3 upstream of the candidate gene LCORL. The highly associated single nucleotide polymorphism BIEC2-808543 (-log(10)P = 8.3) and the adjacent gene LCORL [is] the most promising candidate for body size. We investigated the relative expression levels of LCORL and its two neighbouring genes NCAPG and DCAF16 using quantitative real-time PCR (RT-qPCR). We could demonstrate a significant association of the relative LCORL expression levels with the size of the horses and the BIEC2-808543 genotypes within and across horse breeds. In heterozygous C/T-horses expression levels of LCORL were significantly decreased by 40% and in homozygous C/C-horses by 56% relative to the smaller T/T-horses. Bioinformatic analyses indicated that this SNP T>C mutation is disrupting a putative binding site of the transcription factor TFIID which is important for the transcription process of genes involved in skeletal bone development. Thus, our findings suggest that expression levels of LCORL play a key role for body size within and across horse breeds and regulation of the expression of LCORL is associated with genetic variants of BIEC2-808543."
" the C allele of BIEC2-808543 is presumably the reason for the reduced expression of LCORL in larger sized horses. LCORL, also known as the Mblk1-related protein, shows characteristic motifs of transcription factors and analyses with mouse tissues indicate that it is able to activate transcription."<-So less LCORL equals greater height.
"In human genome-wide scans for adult stature evidences LCORL to be associated with trunk length and hip axis length"
"the correlation between growth of body and hair could be shown in various studies. The Rothmund-Thomson syndrome in human for example is characterized by severe dwarfism combined with an abnormal hair growth. Studies in mice revealed growth retardation in hair length and a retarded rate of body growth caused by the supply of high concentrations of the epidermal growth factor (EGF)"
So we can look at hair growth to find height genes.
"The candidate genes NCAPG and DCAF16 could be eliminated as candidates [for human height], whereas HMGA2 (high mobility group AT-hook 2, ECA6), ZFAT (zinc finger and AT hook domain containing, ECA9), LASP1 (LIM and SH3 protein 1, ECA11) could also possibly involved."
Four loci explain 83% of size variation in the horse.
"Unlike humans, which are naturally reproducing and possess many genetic variants with weak effects on size, we show that horses, like other domestic mammals, carry just a small number of size loci with alleles of large effect. Furthermore, three of our horse size loci contain the LCORL, HMGA2 and ZFAT genes that have previously been found to control human height. The LCORL/NCAPG locus is also implicated in cattle growth and HMGA2 is associated with dog size. Extreme size diversification is a hallmark of domestication."
"the immediately adjacent gene [to LCORL], NCAPG, has been implicated in prenatal growth"
"HMGA2 is an architectural transcription factor that regulates gene expression and directs cellular growth, proliferation and differentiation"
"1% of all human genes are now implicated in contributing to size variation"
Reaching new heights: insights into the genetics of human stature
"The height of an individual is the result of many growth and development processes and greater stature is not just a result of increasing bone length; tissue and organ sizes are usually also proportionately increased."
HMGA2 and GDF5 are two genes listed as highly influencing height.
LCORL along with c4orf30 are in the same gene loci as NCAPG.
Labels:
LCORL
Monday, February 23, 2009
Periosteal Stripping
Stimulation of the Longitudinal Growth of Long Bones by Periosteal Stripping
I couldn't copy and paste from this study. The periosteum was sripped from one epiphyseal plate to another. In the 15 dogs studied periosteal stipping either did not enhance growth or slightly increased growth by usually 1-3%.
Periosteal stripping did cause bone fractures so that could've been part of the growth cause.
Stimulation of Bone Growth by Periosteal Stripping
Another study where I couldn't copy and paste. This study was done on children and an increase in length following periosteal stripping was seen. Age range: 6-15 years old. The mean overgrowth of the periosteal stripped femurs was 0.70cm.
Effects of extensive circumferential periosteal stripping on the microstructure and mechanical properties of the murine femoral cortex.
"Extensive periosteal stripping (PS) is a risk factor for post-radiation pathologic fracture following surgery for extremity soft tissue tumors. The purpose of this study was to determine the effects of PS on bone structure and mechanical properties. Thirty-one skeletally mature mice[12-14 week old female] underwent PS, with circumferential removal of periosteum from an 8-mm segment of the mid-diaphysis of the left femur. Thirty-one control mice underwent sham surgery in which the femur was isolated without manipulation of the periosteum. At 2, 6, 12, or 26 weeks following surgery, the left femora were examined by micro-CT to quantify cortical thickness (CtTh), cross-sectional area (CSA), bone volume (BV), and polar moment of inertia (PMI). Three-point mechanical bend testing was performed and peak load, stiffness, and energy to failure were determined. PS resulted in significantly decreased CtTh, CSA, BV, and PMI at all time points. Peak load, stiffness, and energy to failure were significantly reduced at 2, 6, and 12 weeks. There were no significant differences in mechanical properties at 26 weeks. In this mouse model, extensive circumferential PS resulted in sustained changes in bone structure that were still evident after 6 months, accompanied by reductions in bone strength that persisted for at least 3 months."
"skeletally mature (12-14 week old) female Balb/c mice"<-growth starts to taper off at 12-14 weeks but we can't rule out growth.
Periosteal Stripping versus control.
"Representative 3-dimensional micro-CT reconstructions of femora from mice that underwent periosteal stripping (PS) or sham surgery (sham). Images reflect observations at 12 weeks but similar findings were seen at all four time points." Periosteal stripping mice are visibly longer although this is just a representation.
A decrease in bone volume could be a mechanism for allowing height growth(bone volume was decreased with periosteal stripping
Surgical technique: Lower limb-length equalization by periosteal stripping and periosteal division.
" The procedure consists of total circumferential stripping followed by transverse division of the periosteum at the proximal, middle, and distal shafts of the femur, tibia, and fibula of the shorter limb.
We retrospectively reviewed 11 children with LLD[limb length discrepency] who underwent PSPD[periosteal stripping or periosteal division]. The average LLD was 6 ± 3.8 cm (range, 3-13 cm). The average age of the patients was 9 ± 2.5 years (range, 7-13 years). Orthoroentgenograms were obtained every 6 to 12 months after the surgery. The minimum followup was 24 months (mean, 52 months; range, 24-108 months).
Limb length equalization (LLE) was achieved in eight of 11 patients in an average of 25 ± 17.2 months (range, 12-60 months) and was maintained throughout the followup. LLE was not achieved in three children whose discrepancy was greater than 10 cm, however, PSPD helped decrease the amount of the discrepancy in all three patients. No major complications were observed in any patients.
PSPD stimulates limb length and LLE is achieved in approximately 2 years after the procedure in the majority of the patients. We believe PSPD should be considered as a surgical option for a LLD up to 6 cm."
"Several hypotheses have been proposed to explain the growth stimulation including hypervascularization or release of the mechanical restraint after the periosteal stripping"
[Influence of stripping periosteum on bone formation in guided tissue regeneration].
"10 mm long segmental defects were created in the diaphyses of both radii in 24 New Zealand rabbits. The defect on the control side was covered with a silicone membrane shaped as a tube. On the experimental side, 10 mm periosteum of both sides of the defect was stripped and the defect was covered with a silicone tube. The animals were killed at the 3rd day and the 1st, 2nd, 3rd, 4th, 6th, 10th, 12th weeks. Samples were treated for radiologic and histologic examinations.
(1) periosteal stripping exerts no effect on callus formation in or out of the tube. (2) osteoblasts in the cambium layer of the periosteum come from bone surface, endosteum or the Haversian envelopes. (3) the tissue in the two layers of the periosteum has different origins, the fibrous layer comes from soft tissue and osteoblasts in the cambium from bone surface or Haversian envelopes.
Stripping periosteum has no influence on bone formation in guided tissue regeneration."
Unfortunately I couldn't get this full study.
Bone growth and modeling changes induced by periosteal stripping in the rat.
"In this study, the changes in longitudinal bone growth and metaphyseal modeling induced by mid diaphyseal periosteal stripping in the rat femur were analyzed by means of histomorphometrical techniques. One hundred forty-four male 30-day-old Sprague-Dawley albino rats distributed in 4 groups of 36 were studied: a control group, a sham group, a group with middiaphyseal right femoral periosteal stripping, and a group with a polyethylene ring wrapped around the stripped zone. The animals were euthanized at 1, 2, or 4 weeks from the start of the experiment, after double tetracycline labeling. A statistically significant, albeit small, longitudinal overgrowth of stripped femurs was observed after a latency period of 2 to 4 weeks. The metaphyseal diameters were greater in stripped femurs than nonstripped femurs. This finding was associated with a lower osteoclastic index in the external metaphyseal surface and with a lower bone formation rate in the internal surface of the metaphyseal cortex."
Couldn't get this study either. So stripping periosteum reduces bone resorption on the external metaphsyeal surface and reduces bone formation on the internal metaphyseal surface. So more bone is being formed on the external metaphyseal surface. Would this increase height?
Periosteal stripping in achondroplastic children. Little effect on limb length in 10 cases.
[couldn't get full study]
"We present a prospective study of the results of periosteal stripping and division in 10 achondroplastic children. A single limb (femur and tibia) was operated on and the change in actual length of each bone and the percentage change in growth compared to that of the non-operated limb was measured by scanogram. The mean absolute increase in growth was small, measuring 3 mm for the femur and 2 mm for the tibia. There was no measurable growth difference after 18 months. This method of increasing limb length in achondroplastic children prior to definitive and extensive lengthening procedures is not recommended."
Usually achondroplastic children have elevated levels of FGFR3 decreasing height so maybe periosteal stripping doesn't work if FGFR3 levels are too high or the periosteum is a limiting factor. If the periosteum is limiting your growth removing it will increase height. If something else is limiting the growth then periosteal stripping won't affect height.
"The cause of overgrowth of the long bone due to PSPD include changes in the hemodynamics and the external
force applied to the growth plate. periosteal stripping [causes] a blockage of blood flow
to the cortical bone and increased blood flow to the growth plate"
I couldn't copy and paste from this study. The periosteum was sripped from one epiphyseal plate to another. In the 15 dogs studied periosteal stipping either did not enhance growth or slightly increased growth by usually 1-3%.
Periosteal stripping did cause bone fractures so that could've been part of the growth cause.
Stimulation of Bone Growth by Periosteal Stripping
Another study where I couldn't copy and paste. This study was done on children and an increase in length following periosteal stripping was seen. Age range: 6-15 years old. The mean overgrowth of the periosteal stripped femurs was 0.70cm.
Effects of extensive circumferential periosteal stripping on the microstructure and mechanical properties of the murine femoral cortex.
"Extensive periosteal stripping (PS) is a risk factor for post-radiation pathologic fracture following surgery for extremity soft tissue tumors. The purpose of this study was to determine the effects of PS on bone structure and mechanical properties. Thirty-one skeletally mature mice[12-14 week old female] underwent PS, with circumferential removal of periosteum from an 8-mm segment of the mid-diaphysis of the left femur. Thirty-one control mice underwent sham surgery in which the femur was isolated without manipulation of the periosteum. At 2, 6, 12, or 26 weeks following surgery, the left femora were examined by micro-CT to quantify cortical thickness (CtTh), cross-sectional area (CSA), bone volume (BV), and polar moment of inertia (PMI). Three-point mechanical bend testing was performed and peak load, stiffness, and energy to failure were determined. PS resulted in significantly decreased CtTh, CSA, BV, and PMI at all time points. Peak load, stiffness, and energy to failure were significantly reduced at 2, 6, and 12 weeks. There were no significant differences in mechanical properties at 26 weeks. In this mouse model, extensive circumferential PS resulted in sustained changes in bone structure that were still evident after 6 months, accompanied by reductions in bone strength that persisted for at least 3 months."
"skeletally mature (12-14 week old) female Balb/c mice"<-growth starts to taper off at 12-14 weeks but we can't rule out growth.
Periosteal Stripping versus control.
"Representative 3-dimensional micro-CT reconstructions of femora from mice that underwent periosteal stripping (PS) or sham surgery (sham). Images reflect observations at 12 weeks but similar findings were seen at all four time points." Periosteal stripping mice are visibly longer although this is just a representation.
A decrease in bone volume could be a mechanism for allowing height growth(bone volume was decreased with periosteal stripping
Surgical technique: Lower limb-length equalization by periosteal stripping and periosteal division.
" The procedure consists of total circumferential stripping followed by transverse division of the periosteum at the proximal, middle, and distal shafts of the femur, tibia, and fibula of the shorter limb.
We retrospectively reviewed 11 children with LLD[limb length discrepency] who underwent PSPD[periosteal stripping or periosteal division]. The average LLD was 6 ± 3.8 cm (range, 3-13 cm). The average age of the patients was 9 ± 2.5 years (range, 7-13 years). Orthoroentgenograms were obtained every 6 to 12 months after the surgery. The minimum followup was 24 months (mean, 52 months; range, 24-108 months).
Limb length equalization (LLE) was achieved in eight of 11 patients in an average of 25 ± 17.2 months (range, 12-60 months) and was maintained throughout the followup. LLE was not achieved in three children whose discrepancy was greater than 10 cm, however, PSPD helped decrease the amount of the discrepancy in all three patients. No major complications were observed in any patients.
PSPD stimulates limb length and LLE is achieved in approximately 2 years after the procedure in the majority of the patients. We believe PSPD should be considered as a surgical option for a LLD up to 6 cm."
"Several hypotheses have been proposed to explain the growth stimulation including hypervascularization or release of the mechanical restraint after the periosteal stripping"
[Influence of stripping periosteum on bone formation in guided tissue regeneration].
"10 mm long segmental defects were created in the diaphyses of both radii in 24 New Zealand rabbits. The defect on the control side was covered with a silicone membrane shaped as a tube. On the experimental side, 10 mm periosteum of both sides of the defect was stripped and the defect was covered with a silicone tube. The animals were killed at the 3rd day and the 1st, 2nd, 3rd, 4th, 6th, 10th, 12th weeks. Samples were treated for radiologic and histologic examinations.
(1) periosteal stripping exerts no effect on callus formation in or out of the tube. (2) osteoblasts in the cambium layer of the periosteum come from bone surface, endosteum or the Haversian envelopes. (3) the tissue in the two layers of the periosteum has different origins, the fibrous layer comes from soft tissue and osteoblasts in the cambium from bone surface or Haversian envelopes.
Stripping periosteum has no influence on bone formation in guided tissue regeneration."
Unfortunately I couldn't get this full study.
Bone growth and modeling changes induced by periosteal stripping in the rat.
"In this study, the changes in longitudinal bone growth and metaphyseal modeling induced by mid diaphyseal periosteal stripping in the rat femur were analyzed by means of histomorphometrical techniques. One hundred forty-four male 30-day-old Sprague-Dawley albino rats distributed in 4 groups of 36 were studied: a control group, a sham group, a group with middiaphyseal right femoral periosteal stripping, and a group with a polyethylene ring wrapped around the stripped zone. The animals were euthanized at 1, 2, or 4 weeks from the start of the experiment, after double tetracycline labeling. A statistically significant, albeit small, longitudinal overgrowth of stripped femurs was observed after a latency period of 2 to 4 weeks. The metaphyseal diameters were greater in stripped femurs than nonstripped femurs. This finding was associated with a lower osteoclastic index in the external metaphyseal surface and with a lower bone formation rate in the internal surface of the metaphyseal cortex."
Couldn't get this study either. So stripping periosteum reduces bone resorption on the external metaphsyeal surface and reduces bone formation on the internal metaphyseal surface. So more bone is being formed on the external metaphyseal surface. Would this increase height?
Periosteal stripping in achondroplastic children. Little effect on limb length in 10 cases.
[couldn't get full study]
"We present a prospective study of the results of periosteal stripping and division in 10 achondroplastic children. A single limb (femur and tibia) was operated on and the change in actual length of each bone and the percentage change in growth compared to that of the non-operated limb was measured by scanogram. The mean absolute increase in growth was small, measuring 3 mm for the femur and 2 mm for the tibia. There was no measurable growth difference after 18 months. This method of increasing limb length in achondroplastic children prior to definitive and extensive lengthening procedures is not recommended."
Usually achondroplastic children have elevated levels of FGFR3 decreasing height so maybe periosteal stripping doesn't work if FGFR3 levels are too high or the periosteum is a limiting factor. If the periosteum is limiting your growth removing it will increase height. If something else is limiting the growth then periosteal stripping won't affect height.
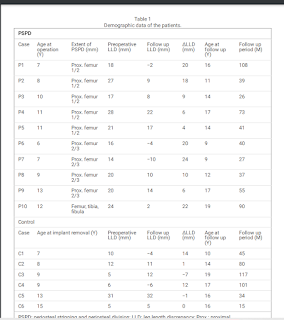
"Periosteal stripping and periosteal division (PSPD) promotes growth of the long bone of children
with leg length discrepancy (LLD). We performed PSPD when LLD was observed at the time of implant removal
surgery after proximal femoral osteotomy for Perthes disease. This study aimed to clarify the efficacy and safety
of PSPD for acquired LLD related to Perthes disease.
Methods: This retrospective study enrolled 10 patients treated with PSPD and 6 control patients who declined the
PSPD for LLD associated with Perthes disease. The lengths of the femur, tibia and entire leg were measured in the
full-length standing radiographs at baseline and final follow-up. Baseline was defined as the time of the last
preoperative observation. LLD and changes in LLD (ΔLLD) were measured. The correlation of ΔLLD with age at
time of surgery, follow-up period, and extent of PSPD was investigated.
Results: Patients’ mean age and LLD at baseline were 9.4 years and 20.5 ± 4.6 mm in the PSPD group and 10.2
years and 11.5 ± 10.0 mm in the control group. With a mean follow-up period of 4.3 years, the PSPD group
showed a mean ΔLLD decrease of 13.9 mm, which was significantly greater than that of the control group at 3.2
mm with a mean follow-up period of 5.4 years. Logistic regression analysis revealed that age at the time of
surgery was a significant factor for obtaining >10 mm ΔLLD with PSPD and the cutoff value by the receiver
operating characteristic curve was 9.6 years (sensitivity, 0.83; specificity 0.83).
Conclusion: PSPD seemed to be a safe and effective surgical option for LLD associated with Perthes disease. The
age at the time of surgery negatively correlated with the amount of LLD correction. Obtaining >10 mm LLD
correction is more likely if the patients are <10 years of age."
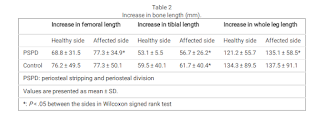
". In the control group, the
affected side also tended to show greater changes than the healthy side, but only the tibia length showed a
significant difference in the amount of change"<-looking at the control group there seems to be mixed results with periosteal stripping but if you look at the table below you see that it does seem to increase leg length.
Labels:
periosteal stripping
Wednesday, February 18, 2009
TAK1
TAK1 regulates LMP1 regulation by TGF-Beta. TAK1 activates MKK3/6.
TAK1 regulates cartilage and joint development via the MAPK and BMP signaling pathways.
"TGF-beta activated kinase 1 (TAK1) is a MAP3K activated by TGF-beta, BMP, and other mitogen-activated protein kinase (MAPK) signaling components. [We delete] Tak1 in chondrocytes (Col2Cre;Tak1(f/f)) and the developing limb mesenchyme (Prx1Cre;Tak1(f/f)). Deletion of Tak1 in chondrocytes resulted in novel embryonic developmental cartilage defects including decreased chondrocyte proliferation, reduced proliferating chondrocyte survival, delayed onset of hypertrophy, reduced Mmp13 expression, and a failure to maintain interzone cells of the elbow joint, which were not observed previously in another Col2Cre;Tak1(f/f) model. Deletion of Tak1 in limb mesenchyme resulted in widespread joint fusions likely owing to the differentiation of interzone cells to the chondrocyte lineage. Loss of Tak1 results in impaired activation of the downstream MAPK target p38, as well as diminished activation of the BMP/SMAD signaling pathway."
"Mouse models lacking the TGF-β type II receptor in chondrocytes (Col2Cre;Tgfbr2f/f) exhibit defects in the postnatal regulation of chondrocyte maturation primarily within the axial skeleton. Deletion of Tgfbr2 in early limb mesenchyme (Prx1Cre;Tgfbr2f/f) resulted in delayed cartilage formation, reduced chondrocyte proliferation, and malformation of joints within the digits."
"Mice lacking BMP receptors 1a and 1b within cartilage (Col2Cre;Bmpr1af/f, Bmpr1b+/−, and Col2Cre;Bmpr1a f/f;Bmpr1b−/−) or lacking canonical BMP targets Smads 1 and 5 within cartilage (Col2Cre;Smad1f/f;Smad5f/f) have reduced chondrocyte proliferation, delayed onset of chondrocyte maturation, reduced proliferating chondrocyte survival, and delayed progression of terminal maturation. Conditional deletion of the BMP ligands, BMP-2 and BMP-4, using the Prx1Cre transgene (Prx1Cre;Bmp2f/f;Bmp4f/f) resulted in shorter and thinner limbs, delayed differentiation and marrow cavity formation, disorganized clearance of hypertrophic chondrocytes, joint fusion of the stylopod and zeugopod elements with little effect on the autopod, and incomplete bone formation. Conventional deletion of another BMP family member, Gdf5, also demonstrated effects on the limb skeleton, resulting in decreased limb length and various joint fusions/abnormalities."
" transgenic mice overexpressing constitutively active MKK6 in chondrocytes are dwarfed, have decreased chondrocyte proliferation, delayed onset of hypertrophic differentiation during embryonic development, and a shortened zone of hypertrophic chondrocytes."
"Treatment of sternal chondrocytes with TAK1 inhibitor TI-2 (LLZ1640-2, 3 µM) resulted in decreased Smad 1/5/8 and p38 phosphorylation"
"loss of Tak1 via Cre-mediated deletion decreased BMP-2-induced phosphorylation of Smads 1/5/8"
TAK1 regulates cartilage and joint development via the MAPK and BMP signaling pathways.
"TGF-beta activated kinase 1 (TAK1) is a MAP3K activated by TGF-beta, BMP, and other mitogen-activated protein kinase (MAPK) signaling components. [We delete] Tak1 in chondrocytes (Col2Cre;Tak1(f/f)) and the developing limb mesenchyme (Prx1Cre;Tak1(f/f)). Deletion of Tak1 in chondrocytes resulted in novel embryonic developmental cartilage defects including decreased chondrocyte proliferation, reduced proliferating chondrocyte survival, delayed onset of hypertrophy, reduced Mmp13 expression, and a failure to maintain interzone cells of the elbow joint, which were not observed previously in another Col2Cre;Tak1(f/f) model. Deletion of Tak1 in limb mesenchyme resulted in widespread joint fusions likely owing to the differentiation of interzone cells to the chondrocyte lineage. Loss of Tak1 results in impaired activation of the downstream MAPK target p38, as well as diminished activation of the BMP/SMAD signaling pathway."
"Mouse models lacking the TGF-β type II receptor in chondrocytes (Col2Cre;Tgfbr2f/f) exhibit defects in the postnatal regulation of chondrocyte maturation primarily within the axial skeleton. Deletion of Tgfbr2 in early limb mesenchyme (Prx1Cre;Tgfbr2f/f) resulted in delayed cartilage formation, reduced chondrocyte proliferation, and malformation of joints within the digits."
"Mice lacking BMP receptors 1a and 1b within cartilage (Col2Cre;Bmpr1af/f, Bmpr1b+/−, and Col2Cre;Bmpr1a f/f;Bmpr1b−/−) or lacking canonical BMP targets Smads 1 and 5 within cartilage (Col2Cre;Smad1f/f;Smad5f/f) have reduced chondrocyte proliferation, delayed onset of chondrocyte maturation, reduced proliferating chondrocyte survival, and delayed progression of terminal maturation. Conditional deletion of the BMP ligands, BMP-2 and BMP-4, using the Prx1Cre transgene (Prx1Cre;Bmp2f/f;Bmp4f/f) resulted in shorter and thinner limbs, delayed differentiation and marrow cavity formation, disorganized clearance of hypertrophic chondrocytes, joint fusion of the stylopod and zeugopod elements with little effect on the autopod, and incomplete bone formation. Conventional deletion of another BMP family member, Gdf5, also demonstrated effects on the limb skeleton, resulting in decreased limb length and various joint fusions/abnormalities."
" transgenic mice overexpressing constitutively active MKK6 in chondrocytes are dwarfed, have decreased chondrocyte proliferation, delayed onset of hypertrophic differentiation during embryonic development, and a shortened zone of hypertrophic chondrocytes."
"Treatment of sternal chondrocytes with TAK1 inhibitor TI-2 (LLZ1640-2, 3 µM) resulted in decreased Smad 1/5/8 and p38 phosphorylation"
"loss of Tak1 via Cre-mediated deletion decreased BMP-2-induced phosphorylation of Smads 1/5/8"
Labels:
TAK1
Cadmium
I think this is a Cadmium supplement: Boiron Cadmium Sulphuricum 6C 75 6c pellets .
.
Cadmium can cause apoptosis of mesenchymal progenitors and inhibit chondrogenesis.
Effects of sub-toxic Cadmium concentrations on bone gene expression program: results of an in vitro study.
"Exposure to the heavy metal Cadmium (Cd) affects human health. This study investigated the effects of exposure to a single, or multiple, sub-toxic Cd concentrations on sub-confluent and confluent human osteoblast growth. RT-PCR quantified gene expression of type I collagen, metalloprotease (MMP13), runt-related transcription factor-2 (RUNX2), osterix, osteocalcin, osteonectin, alkaline phosphatase, integrins and bone sialoprotein (BSP). Expression of fibroblast growth factors 1 and 2 (FGF1, FGF2), transforming growth factor-beta(3) (TGFbeta(3)) and bone morphogenetic protein-2 (BMP2) were also evaluated to determine whether Cd-related effects were mediated by an imbalance in expression. Depending on osteoblast concentration and maturation stages, Cd inhibited or stimulated cell growth, decreased type I collagen, increased MMP13, FGF1 and BMP2{BMP2 and MMP13 genes can affect height} gene expression and stimulated the mineralization process only in continuously exposed cultures. In vivo, acute or chronic exposure to sub-toxic Cd concentrations may affect bone formation differently and support the hypothesis that Cd-induced bone disorders may involve downstream changes in growth factor expression."
The osteoblasts were all from old individuals(50+).
"On day 22, untreated and Cd-treated osteoblasts, cultured according to protocol B, grew uniform confluent sheets of fibroblast-like and cubical cells, with nodular structures. These were apparently the result of coalescing aggregates under the inverted microscope"<-fibroblast cells can be precursors to chondrocyte cells.
"protocol B cultures were maintained in vitro for 22 days and exposed to Cd for the last 3 days"
Protocol B reduced type I collagen, Runx2, BSP, and osterix expression while increasing MMP13 expression so it was the most chondrogenic. The less chondrogenic protocols involved adding cadmium earlier or continuously adding cadmium to the medium.
However, Protocol B also decreased the expression of the pro-chondrogenic genes BMP-2 and TGF-Beta3. Only continuous administration of Cadmium increased BMP-2 levels.
" Cd had an inhibitory effect on cell number when added to a low density cellular sheet, while 0.1 μM Cd-stimulated cell number in 12-day sub-confluent and in 22-day confluent cultures."<-Higher density cells are more chondrogenic.
Cadmium can cause apoptosis of mesenchymal progenitors and inhibit chondrogenesis.
Effects of sub-toxic Cadmium concentrations on bone gene expression program: results of an in vitro study.
"Exposure to the heavy metal Cadmium (Cd) affects human health. This study investigated the effects of exposure to a single, or multiple, sub-toxic Cd concentrations on sub-confluent and confluent human osteoblast growth. RT-PCR quantified gene expression of type I collagen, metalloprotease (MMP13), runt-related transcription factor-2 (RUNX2), osterix, osteocalcin, osteonectin, alkaline phosphatase, integrins and bone sialoprotein (BSP). Expression of fibroblast growth factors 1 and 2 (FGF1, FGF2), transforming growth factor-beta(3) (TGFbeta(3)) and bone morphogenetic protein-2 (BMP2) were also evaluated to determine whether Cd-related effects were mediated by an imbalance in expression. Depending on osteoblast concentration and maturation stages, Cd inhibited or stimulated cell growth, decreased type I collagen, increased MMP13, FGF1 and BMP2{BMP2 and MMP13 genes can affect height} gene expression and stimulated the mineralization process only in continuously exposed cultures. In vivo, acute or chronic exposure to sub-toxic Cd concentrations may affect bone formation differently and support the hypothesis that Cd-induced bone disorders may involve downstream changes in growth factor expression."
The osteoblasts were all from old individuals(50+).
"On day 22, untreated and Cd-treated osteoblasts, cultured according to protocol B, grew uniform confluent sheets of fibroblast-like and cubical cells, with nodular structures. These were apparently the result of coalescing aggregates under the inverted microscope"<-fibroblast cells can be precursors to chondrocyte cells.
"protocol B cultures were maintained in vitro for 22 days and exposed to Cd for the last 3 days"
Protocol B reduced type I collagen, Runx2, BSP, and osterix expression while increasing MMP13 expression so it was the most chondrogenic. The less chondrogenic protocols involved adding cadmium earlier or continuously adding cadmium to the medium.
However, Protocol B also decreased the expression of the pro-chondrogenic genes BMP-2 and TGF-Beta3. Only continuous administration of Cadmium increased BMP-2 levels.
" Cd had an inhibitory effect on cell number when added to a low density cellular sheet, while 0.1 μM Cd-stimulated cell number in 12-day sub-confluent and in 22-day confluent cultures."<-Higher density cells are more chondrogenic.
Labels:
cadmium
Sunday, February 8, 2009
Itm2a
Parathyroid Hormone increases Itm2a. LSJL upregulates a form of Itm2a. Itm2a is a gene associated with height growth. There is a patent for an antibody to inhibit Itm2a.
Enhanced ITM2A expression inhibits chondrogenic differentiation of mesenchymal stem cells.
Enhanced ITM2A expression inhibits chondrogenic differentiation of mesenchymal stem cells.
"The aim of this work was to characterise ASC in comparison to MSC in order to identify genes which may be involved in mechanisms causing the altered chondrogenic potential of ASC. Representational difference analysis was used to identify genes with higher expression in undifferentiated ASC than in MSC. Integral membrane protein 2A (ITM2A) was higher expressed in expanded ASC than in MSC in a donor-independent manner. During early chondrogenic differentiation in spheroid cultures ITM2A levels remained low in MSC and a transient down-regulation occurred in ASC correlating with successful chondrogenesis. Persisting ITM2A levels were found in non-differentiating ASC. Consistent with this finding, forced expression of ITM2A in the mouse mesenchymal stem cell line C3H10T1/2 prevented chondrogenic induction. ITM2A may in early stages of differentiation be associated with an inhibition of the initiation of chondrogenesis and elevated expression of ITM2A in ASC may therefore be linked to the poorer chondrogenic differentiation potential of these cells."
Even though Itm2a may inhibit chondrogenesis it may still be a marker for early endochondral ossification and may be evidence that LSJL can induce new growth plates.
"COL2A1 mRNA expression was not induced in the ITM2A over-expressing cells"
"Strong ITM2A expression has been shown in chondrocytes of the resting zone of the murine growth plate."
ITM2A is much more strongly expressed in chonondrogenic differentiation MSCs and chondrocytes than osteoblasts and adipo- and osteo- differentiating MSCs.
Genes more strongly expressed in ASC than MSCs also upregulated in LSJL:
MMP3
ITM2A
GALNTL1
Collagenase-3 (MMP-13) and integral membrane protein 2a (Itm2a) are marker genes of chondrogenic/osteoblastic cells in bone formation: sequential temporal, and spatial expression of Itm2a, alkaline phosphatase, MMP-13, and osteocalcin in the mouse.
" we compared the expression pattern of the recently cloned Itm2a and MMP-13 (collagenase-3) genes with that of established marker genes for bone formation, such as alkaline phosphatase (ALP), osteocalcin (OC), and collagen type X, during endochondral and intramembranous ossification. During embryonic development expression of Itm2a and ALP was detectable at midgestation (11.5 days postcoitum [dpc]) and increased up to 16.5 dpc. MMP-13 and OC expression started at 14.5 dpc and 16.5 dpc, respectively. This temporal expression was reflected in the spatial distribution of these markers in the growth plate of long bones. In areas undergoing endochondral ossification Itm2a expression was found in chondrocytes of the resting and the proliferating zones. Expression of ALP and MMP-13 are mutually exclusive: ALP transcripts were found only in collagen type X positive hypertrophic chondrocytes of the upper zone. MMP-13 expression was restricted to chondrocytes of the lower zone of hypertrophic cartilage also expressing collagen type X. In osteoblasts involved in endochondral and intramembranous ossification Itm2a was not present. ALP, MMP-13, and OC were mutually exclusively expressed in these cells suggesting a differentiation-dependent sequential expression of ALP, MMP-13, and OC. The identification of the continuum of sequential expression of Itm2a, ALP, MMP-13, and OC will now allow us to establish a series of marker genes that are highly suitable to characterize bone cells during chondrocytic and osteoblastic differentiation in vivo."
"Hybridization with the Itm2a probe showed signals in the periosteum and perichondrium of the tibia and in the distal part of the growth plate. Cells of the mature bone did not express Itm2a at significant levels. Chondrocytes of the resting zone in the growth plate showed strong signals of Itm2a expression. In the proliferative zone, laterally located chondrocytes expressed Itm2a"
Labels:
Itm2a
Lef1
Overexpression of Lef1 diminishes the effects of Beta-Catenin on Notch1. Lef1 is more strongly expressed in growth plate cartilage than articular cartilage.
Looping Mediated Interaction between the Promoter and 3′ UTR Regulates Type II Collagen Expression in Chondrocytes
"Type II collagen is the major component of articular cartilage and is mainly synthesized by chondrocytes. Repeated sub-culturing of primary chondrocytes leads to reduction of type II collagen gene (Col2a1) expression, which mimics the process of chondrocyte dedifferentiation. We have investigated the crosstalk between cis-acting DNA element and transcription factor on Col2a1 expression in primary chondrocytes. Bioinformatic analysis revealed the potential regulatory regions in the Col2a1 genomic locus. Among them, promoter and 3′ untranslated region (UTR) showed highly accessible chromatin architecture with enriched recruitment of active chromatin markers in primary chondrocytes. 3′ UTR has a potent enhancer function which recruits Lef1 (Lymphoid enhancer binding factor 1) transcription factor, leading to juxtaposition of the 3′ UTR with the promoter through gene looping resulting in up-regulation of Col2a1 gene transcription. Knock-down of endogenous Lef1 level significantly reduced the gene looping and subsequently down-regulated Col2a1 expression. However, these regulatory loci become inaccessible due to condensed chromatin architecture as chondrocytes dedifferentiate which was accompanied by a reduction of gene looping and down-regulation of Col2a1 expression. Lef1 mediated looping between promoter and 3′ UTR under the permissive chromatin architecture upregulates Col2a1 expression in primary chondrocytes."
"Lef1 over-expression significantly increased Col2a1 expression"
"Expression of TCF-3, TCF-4 and Lef1 have been reported in the cartilage. Upregulated TCF-1 expression is confined to the prehypertrophic chondrocytes and in the surrounding perichondrium. TCF-3 can be detected in the whole cartilage, while hypertrophic chondrocytes in Col2a1-ICAT transgenic mice only express TCF-4"
" β-catenin is essential for the early stages of cartilage growth and development whereas over-expression of β-catenin in the adult cartilage is accompanied by cartilage destruction. Inhibition of β-catenin signaling in chondrocytes resulted in significant reduction of expression of Lef/TCF family members such as Lef1, TCF-3 and TCF-4 and showed defects in post-natal cartilage development and delayed chondrogenesis which is accompanied by significant decrease in the cartilage marker gene Col2a1"
"Absence of Lef1 leads to increased apoptosis of chondrocytes and similar effect is also found upon Col2a1 deficiency. Lef1 suppressed cells reduced expression of Col11a1{up}"
"Col11a1, a heterotrimer of alpha 1 (XI) collagen (Col11a1), alpha 2 (XI) collagen (Col11a2) and alpha 1 (II) collagen, is mainly expressed in the articular cartilage and vitreous fluid of the eye, and is responsible for the proper type II collagen fibril formation"
"Mutation of Type XI{up} collagen resulted in accumulation of degraded type II collagen in articular cartilage"
Since LSJL gene expression was taken at 1 hour after last loading, it's possible that Lef1 was upregulated sooner than that and the residual effects are taken at one hour.
Labels:
Lef1
Wednesday, February 4, 2009
SIRT1
SIRT1 regulates differentiation of mesenchymal stem cells by deacetylating β-catenin.
"[We created] MSC specific SIRT1 knock-out (MSCKO) mice. Aged MSCKO mice (2.2 years old) showed defects in tissues derived from MSCs; i.e. a reduction in subcutaneous fat, cortical bone thickness and trabecular volume. Young mice showed related but less pronounced effects. MSCs isolated from MSCKO mice showed reduced differentiation towards osteoblasts and chondrocytes in vitro, but no difference in proliferation or apoptosis. Expression of β-catenin targets important for differentiation was reduced in MSCKO cells. Moreover, while β-catenin itself (T41A mutant resistant to cytosolic turnover) accumulated in the nuclei of wild-type MSCs, it was unable to do so in MSCKO cells. However, mutating K49R or K345R in β-catenin to mimic deacetylation restored nuclear localization and differentiation potential in MSCKO cells. SIRT1 deacetylates β-catenin to promote its accumulation in the nucleus leading to transcription of genes for MSC differentiation."
"SirT1 heterozygotes show increased osteoarthritis and increased levels of chondrocyte apoptosis in cartilage"
"Cartilage of the long bones in MSCKO mice showed no morphological change"<-so maybe early expression of Beta-Catenin does not inhibit longitudinal bone growth
"when MSCs differentiate, levels of SIRT1 decrease, and acetylation of β-catenin increases"
I couldn't access the supplementary data files and it looked like there was some good info in there.
Set7/9 impacts COL2A1 expression through binding and repression of SirT1 histone deacetylation.
"COL2A1 gene expression is positively regulated by the NAD-dependent protein deacetylase SirT1, through its ability to bind chromatin regions of the COL2A1 promoter and enhancer. Human chondrocytes were encapsulated in three-dimensional (3D) alginate beads where they exhibited up-regulated COL2A1 mRNA expression and increased levels of SirT1 occupancy on the promoter and enhancer regions, when compared to monolayer controls. Chromatin immunoprecipitation (ChIP) analyses of 3D cultures showed augmentated levels of the DNA-binding transcription factor SP1, and the histone methyltransferase Set7/9, on the COL2A1 promoter site. ChIP reChIP assays revealed that SirT1 and Set7/9 form a protein complex on the COL2A1 promoter region of 3D-cultured chondrocytes, which also demonstrated elevated trimethylated lysine 4 on histone 3 (3MeH3K4), a hallmark of Set7/9 methyltransferase activity. Advanced passaging of chondrocytes yielded a decrease in 3MeH3K4 and Set7/9 levels on the COL2A1 promoter and reduced COL2A1 expression, suggesting that the SirT1/Set7/9 complex is preferentially formed on the COL2A1 promoter and required for gene activation. Interestingly, despite SirT1 occupancy, its deacetylation targets (i.e H3K9/14 and H4K16) were found acetylated on the COL2A1 promoter of 3D-cultured chondrocytes. A possible explanation for this phenotype is the enrichment of the histone acetyltransferases P300 and GCN5 on the COL2A1 promoter of3 D-cultured chondrocytes. Our study indicates that Set7/9 prevents the histone deacetylase activity of SirT1, potentiating euchromatin formation on the promoter site of COL2A1 and resulting in morphology-dependent COL2A1 gene transactivation."
"human chondrocyte cell lines stably overexpressing SirT1 displayed increased expression of COL2A1 mRNA and demonstrate enriched occupancy of SirT1, Sox9 and PGC1α on the gene enhancer region. Similarly, TNFα-stimulated human chondrocytes, which possessed reduced COL2A1 mRNA expression, displayed an inactive SirT1 cleaved variant (75SirT1) that yielded reduced occupancy of PGC1α and Sox9 on the COL2A1 enhancer region"
"chondrocyte passaging reduces cellular SirT1 activity, which is likely a result of increased inhibitory DBC1."
"SirT1 causes Set7/9 association on the COL2A1 promoter, leading to increased levels of 3MeH3K4."
Set7/9 impacts COL2A1 expression through binding and repression of SirT1 histone deacetylation.
"COL2A1 gene expression is positively regulated by the NAD-dependent protein deacetylase SirT1, through its ability to bind chromatin regions of the COL2A1 promoter and enhancer. Human chondrocytes were encapsulated in three-dimensional (3D) alginate beads where they exhibited up-regulated COL2A1 mRNA expression and increased levels of SirT1 occupancy on the promoter and enhancer regions, when compared to monolayer controls. Chromatin immunoprecipitation (ChIP) analyses of 3D cultures showed augmentated levels of the DNA-binding transcription factor SP1, and the histone methyltransferase Set7/9, on the COL2A1 promoter site. ChIP reChIP assays revealed that SirT1 and Set7/9 form a protein complex on the COL2A1 promoter region of 3D-cultured chondrocytes, which also demonstrated elevated trimethylated lysine 4 on histone 3 (3MeH3K4), a hallmark of Set7/9 methyltransferase activity. Advanced passaging of chondrocytes yielded a decrease in 3MeH3K4 and Set7/9 levels on the COL2A1 promoter and reduced COL2A1 expression, suggesting that the SirT1/Set7/9 complex is preferentially formed on the COL2A1 promoter and required for gene activation. Interestingly, despite SirT1 occupancy, its deacetylation targets (i.e H3K9/14 and H4K16) were found acetylated on the COL2A1 promoter of 3D-cultured chondrocytes. A possible explanation for this phenotype is the enrichment of the histone acetyltransferases P300 and GCN5 on the COL2A1 promoter of3 D-cultured chondrocytes. Our study indicates that Set7/9 prevents the histone deacetylase activity of SirT1, potentiating euchromatin formation on the promoter site of COL2A1 and resulting in morphology-dependent COL2A1 gene transactivation."
"human chondrocyte cell lines stably overexpressing SirT1 displayed increased expression of COL2A1 mRNA and demonstrate enriched occupancy of SirT1, Sox9 and PGC1α on the gene enhancer region. Similarly, TNFα-stimulated human chondrocytes, which possessed reduced COL2A1 mRNA expression, displayed an inactive SirT1 cleaved variant (75SirT1) that yielded reduced occupancy of PGC1α and Sox9 on the COL2A1 enhancer region"
"chondrocyte passaging reduces cellular SirT1 activity, which is likely a result of increased inhibitory DBC1."
"SirT1 causes Set7/9 association on the COL2A1 promoter, leading to increased levels of 3MeH3K4."
Labels:
SIRT1
Subscribe to:
Posts (Atom)